With an upcoming move to the DC area, I'm keenly interested in figuring out where a whole new slate of species can be found. I'm also a bit sad that my new job won't allow me to use R, which I just spent the past several years learning!
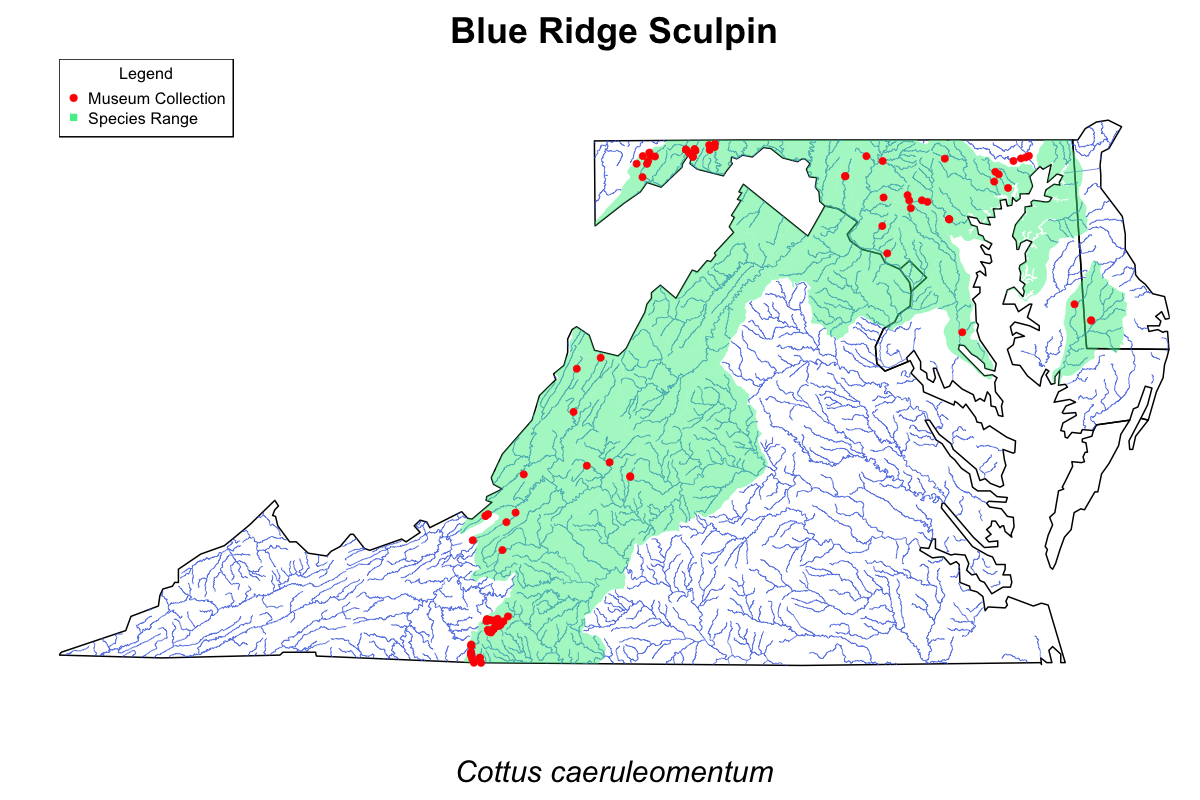
So, to keep using R, and to help myself understand fish distribution, I've started to make a series of maps combining museum collections at http://www.fishnet2.net and watershed distribution maps from NatureServe.
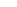
I have a set of ~330 species (mostly freshwater, some saltwater, some from Chesapeake Bay...~70 completed so far) that I'll be making maps for (example attached). Although these are mostly just for my own personal use (I hope to add my own points to the maps as I catch/see various species....and I'm sure there are more accurate maps out there!) I'd be happy to eventually share the whole set online or as a PDF if anyone is interested.
Also, if anyone has any suggestions for the map style or additional sources for collection points, I'm all ears!
References for the attached map:
NatureServe. 2010. Digital Distribution Maps of the Freshwater Fishes in the Conterminous United States. Version 3.0. Arlington, VA. U.S.A.
Fish data used in this study obtained from Mississippi Museum of Natural Science, North Carolina State Museum of Natural Sciences, Ohio State University - Fish Division, University of Alabama Ichthyological Collection, UF, USNM, (Accessed through the Fishnet2 Portal, www.fishnet2.net, 2015-06-12).